Introduction¶
Existing fitting modules are not very pythonic in their API and can be
difficult for humans to use. This project aims to marry the power of
scipy.optimize with the readability of sympy to create a highly
readable and easy to use fitting package which works for projects of any scale.
symfit makes it extremely easy to provide guesses for your parameters
and to bound them to a certain range:
a = Parameter('a', 1.0, min=0.0, max=5.0)
To define models to fit to:
x = Variable('x')
A = Parameter('A')
sig = Parameter('sig', 1.0, min=0.0, max=5.0)
x0 = Parameter('x0', 1.0, min=0.0)
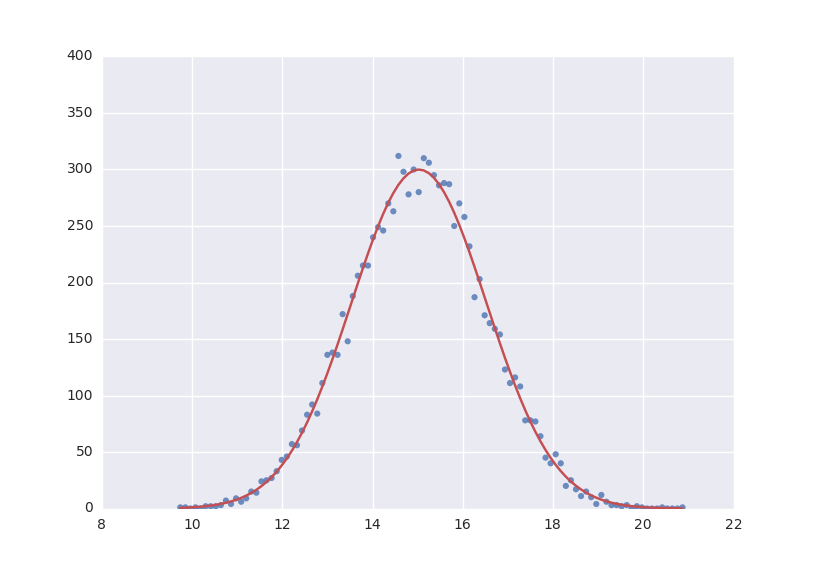
# Gaussian distrubution
model = A * exp(-(x - x0)**2/(2 * sig**2))
And finally, to execute the fit:
fit = Fit(model, xdata, ydata)
fit_result = fit.execute()
And to evaluate the model using the best fit parameters:
y = model(x=xdata, **fit_result.params)
As your models become more complicated, symfit really comes into it’s
own. For example, vector valued functions are both easy to define and beautiful
to look at:
model = {
y_1: x**2,
y_2: 2*x
}
And constrained maximization has never been this easy:
x, y = parameters('x, y')
model = 2*x*y + 2*x - x**2 -2*y**2
constraints = [
Eq(x**3 - y, 0), # Eq: ==
Ge(y - 1, 0), # Ge: >=
]
fit = Fit(- model, constraints=constraints)
fit_result = fit.execute()
Technical Reasons¶
On a more technical note, this symbolic approach turns out to have great
technical advantages over using scipy directly. In order to fit, the algorithm
needs the Jacobian: a matrix containing the derivatives of your model in it’s
parameters. Because of the symbolic nature of symfit, this is determined
for you on the fly, saving you the trouble of having to determine the
derivatives yourself. Furthermore, having this Jacobian allows good estimation
of the errors in your parameters, something scipy does
not always succeed in.